A debate about which language is better suited for Datascience, R or Python, can set off diehard fans of these languages into a tizzy. This post tries to look at some of the different similarities and similar differences between these languages. To a large extent the ease or difficulty in learning R or Python is subjective. I have heard that R has a steeper learning curve than Python and also vice versa. This probably depends on the degree of familiarity with the languuge To a large extent both R an Python do the same thing in just slightly different ways and syntaxes. The ease or the difficulty in the R/Python construct’s largely is in the ‘eyes of the beholder’ nay, programmer’ we could say. I include my own experience with the languages below.
Check out my compact and minimal book “Practical Machine Learning with R and Python:Third edition- Machine Learning in stereo” available in Amazon in paperback($12.99) and kindle($8.99) versions. My book includes implementations of key ML algorithms and associated measures and metrics. The book is ideal for anybody who is familiar with the concepts and would like a quick reference to the different ML algorithms that can be applied to problems and how to select the best model. Pick your copy today!!

1. R data types
R has the following data types
- Character
- Integer
- Numeric
- Logical
- Complex
- Raw
Python has several data types
- Int
- float
- Long
- Complex and so on
2. R Vector vs Python List
A common data type in R is the vector. Python has a similar data type, the list
# R vectors
a<-c(4,5,1,3,4,5)
print(a[3])## [1] 1print(a[3:4]) # R does not always need the explicit print. ## [1] 1 3#R type of variable
print(class(a))## [1] "numeric"# Length of a
print(length(a))## [1] 6# Python lists
a=[4,5,1,3,4,5] #
print(a[2]) # Some python IDEs require the explicit print
print(a[2:5])
print(type(a))
# Length of a
print(len(a))## 1
## [1, 3, 4]
##
## 62a. Other data types – Python
Python also has certain other data types like the tuple, dictionary etc as shown below. R does not have as many of the data types, nevertheless we can do everything that Python does in R
# Python tuple
b = (4,5,7,8)
print(b)
#Python dictionary
c={'name':'Ganesh','age':54,'Work':'Professional'}
print(c)
#Print type of variable c
## (4, 5, 7, 8)
## {'name': 'Ganesh', 'age': 54, 'Work': 'Professional'}2.Type of Variable
To know the type of the variable in R we use ‘class’, In Python the corresponding command is ‘type’
#R - Type of variable
a<-c(4,5,1,3,4,5)
print(class(a))## [1] "numeric"#Python - Print type of tuple a
a=[4,5,1,3,4,5]
print(type(a))
b=(4,3,"the",2)
print(type(b))##
## 3. Length
To know length in R, use length()
#R - Length of vector
# Length of a
a<-c(4,5,1,3,4,5)
print(length(a))## [1] 6To know the length of a list,tuple or dict we can use len()
# Python - Length of list , tuple etc
# Length of a
a=[4,5,1,3,4,5]
print(len(a))
# Length of b
b = (4,5,7,8)
print(len(b))
## 6
## 44. Accessing help
To access help in R we use the ‘?’ or the ‘help’ function
#R - Help - To be done in R console or RStudio
#?sapply
#help(sapply)Help in python on any topic involves
#Python help - This can be done on a (I)Python console
#help(len)
#?len5. Subsetting
The key difference between R and Python with regards to subsetting is that in R the index starts at 1. In Python it starts at 0, much like C,C++ or Java To subset a vector in R we use
#R - Subset
a<-c(4,5,1,3,4,8,12,18,1)
print(a[3])## [1] 1# To print a range or a slice. Print from the 3rd to the 5th element
print(a[3:6])## [1] 1 3 4 8Python also uses indices. The difference in Python is that the index starts from 0/
#Python - Subset
a=[4,5,1,3,4,8,12,18,1]
# Print the 4th element (starts from 0)
print(a[3])
# Print a slice from 4 to 6th element
print(a[3:6])## 3
## [3, 4, 8]6. Operations on vectors in R and operation on lists in Python
In R we can do many operations on vectors for e.g. element by element addition, subtraction, exponentation,product etc. as show
#R - Operations on vectors
a<- c(5,2,3,1,7)
b<- c(1,5,4,6,8)
#Element wise Addition
print(a+b)## [1] 6 7 7 7 15#Element wise subtraction
print(a-b)## [1] 4 -3 -1 -5 -1#Element wise product
print(a*b)## [1] 5 10 12 6 56# Exponentiating the elements of a vector
print(a^2)## [1] 25 4 9 1 49In Python to do this on lists we need to use the ‘map’ and the ‘lambda’ function as follows
# Python - Operations on list
a =[5,2,3,1,7]
b =[1,5,4,6,8]
#Element wise addition with map & lambda
print(list(map(lambda x,y: x+y,a,b)))
#Element wise subtraction
print(list(map(lambda x,y: x-y,a,b)))
#Element wise product
print(list(map(lambda x,y: x*y,a,b)))
# Exponentiating the elements of a list
print(list(map(lambda x: x**2,a)))
## [6, 7, 7, 7, 15]
## [4, -3, -1, -5, -1]
## [5, 10, 12, 6, 56]
## [25, 4, 9, 1, 49]However if we create ndarrays from lists then we can do the element wise addition,subtraction,product, etc. like R. Numpy is really a powerful module with many, many functions for matrix manipulations
import numpy as np
a =[5,2,3,1,7]
b =[1,5,4,6,8]
a=np.array(a)
b=np.array(b)
#Element wise addition
print(a+b)
#Element wise subtraction
print(a-b)
#Element wise product
print(a*b)
# Exponentiating the elements of a list
print(a**2)
## [ 6 7 7 7 15]
## [ 4 -3 -1 -5 -1]
## [ 5 10 12 6 56]
## [25 4 9 1 49]7. Getting the index of element
To determine the index of an element which satisifies a specific logical condition in R use ‘which’. In the code below the index of element which is equal to 1 is 4
# R - Which
a<- c(5,2,3,1,7)
print(which(a == 1))## [1] 4In Python array we can use np.where to get the same effect. The index will be 3 as the index starts from 0
# Python - np.where
import numpy as np
a =[5,2,3,1,7]
a=np.array(a)
print(np.where(a==1))## (array([3], dtype=int64),)8. Data frames
R, by default comes with a set of in-built datasets. There are some datasets which come with the SkiKit- Learn package
# R
# To check built datasets use
#data() - In R console or in R Studio
#iris - Don't print to consoleWe can use the in-built data sets that come with Scikit package
#Python
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
# This creates a Sklearn bunch
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)9. Working with dataframes
With R you can work with dataframes directly. For more complex dataframe operations in R there are convenient packages like dplyr, reshape2 etc. For Python we need to use the Pandas package. Pandas is quite comprehensive in the list of things we can do with data frames The most common operations on a dataframe are
- Check the size of the dataframe
- Take a look at the top 5 or bottom 5 rows of dataframe
- Check the content of the dataframe
a.Size
In R use dim()
#R - Size
dim(iris)## [1] 150 5For Python use .shape
#Python - size
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
iris.shapeb. Top & bottom 5 rows of dataframe
To know the top and bottom rows of a data frame we use head() & tail as shown below for R and Python
#R
head(iris,5)## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 1 5.1 3.5 1.4 0.2 setosa
## 2 4.9 3.0 1.4 0.2 setosa
## 3 4.7 3.2 1.3 0.2 setosa
## 4 4.6 3.1 1.5 0.2 setosa
## 5 5.0 3.6 1.4 0.2 setosatail(iris,5)## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 146 6.7 3.0 5.2 2.3 virginica
## 147 6.3 2.5 5.0 1.9 virginica
## 148 6.5 3.0 5.2 2.0 virginica
## 149 6.2 3.4 5.4 2.3 virginica
## 150 5.9 3.0 5.1 1.8 virginica#Python
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
print(iris.head(5))
print(iris.tail(5))## sepal length (cm) sepal width (cm) petal length (cm) petal width (cm)
## 0 5.1 3.5 1.4 0.2
## 1 4.9 3.0 1.4 0.2
## 2 4.7 3.2 1.3 0.2
## 3 4.6 3.1 1.5 0.2
## 4 5.0 3.6 1.4 0.2
## sepal length (cm) sepal width (cm) petal length (cm) petal width (cm)
## 145 6.7 3.0 5.2 2.3
## 146 6.3 2.5 5.0 1.9
## 147 6.5 3.0 5.2 2.0
## 148 6.2 3.4 5.4 2.3
## 149 5.9 3.0 5.1 1.8c. Check the content of the dataframe
#R
summary(iris)## Sepal.Length Sepal.Width Petal.Length Petal.Width
## Min. :4.300 Min. :2.000 Min. :1.000 Min. :0.100
## 1st Qu.:5.100 1st Qu.:2.800 1st Qu.:1.600 1st Qu.:0.300
## Median :5.800 Median :3.000 Median :4.350 Median :1.300
## Mean :5.843 Mean :3.057 Mean :3.758 Mean :1.199
## 3rd Qu.:6.400 3rd Qu.:3.300 3rd Qu.:5.100 3rd Qu.:1.800
## Max. :7.900 Max. :4.400 Max. :6.900 Max. :2.500
## Species
## setosa :50
## versicolor:50
## virginica :50
##
##
## str(iris)## 'data.frame': 150 obs. of 5 variables:
## $ Sepal.Length: num 5.1 4.9 4.7 4.6 5 5.4 4.6 5 4.4 4.9 ...
## $ Sepal.Width : num 3.5 3 3.2 3.1 3.6 3.9 3.4 3.4 2.9 3.1 ...
## $ Petal.Length: num 1.4 1.4 1.3 1.5 1.4 1.7 1.4 1.5 1.4 1.5 ...
## $ Petal.Width : num 0.2 0.2 0.2 0.2 0.2 0.4 0.3 0.2 0.2 0.1 ...
## $ Species : Factor w/ 3 levels "setosa","versicolor",..: 1 1 1 1 1 1 1 1 1 1 ...#Python
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
print(iris.info())##
## RangeIndex: 150 entries, 0 to 149
## Data columns (total 4 columns):
## sepal length (cm) 150 non-null float64
## sepal width (cm) 150 non-null float64
## petal length (cm) 150 non-null float64
## petal width (cm) 150 non-null float64
## dtypes: float64(4)
## memory usage: 4.8 KB
## Noned. Check column names
#R
names(iris)## [1] "Sepal.Length" "Sepal.Width" "Petal.Length" "Petal.Width"
## [5] "Species"colnames(iris)## [1] "Sepal.Length" "Sepal.Width" "Petal.Length" "Petal.Width"
## [5] "Species"#Python
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
#Get column names
print(iris.columns)## Index(['sepal length (cm)', 'sepal width (cm)', 'petal length (cm)',
## 'petal width (cm)'],
## dtype='object')e. Rename columns
In R we can assign a vector to column names
#R
colnames(iris) <- c("lengthOfSepal","widthOfSepal","lengthOfPetal","widthOfPetal","Species")
colnames(iris)## [1] "lengthOfSepal" "widthOfSepal" "lengthOfPetal" "widthOfPetal"
## [5] "Species"In Python we can assign a list to s.columns
#Python
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
iris.columns = ["lengthOfSepal","widthOfSepal","lengthOfPetal","widthOfPetal"]
print(iris.columns)## Index(['lengthOfSepal', 'widthOfSepal', 'lengthOfPetal', 'widthOfPetal'], dtype='object')f.Details of dataframe
#Python
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
print(iris.info())##
## RangeIndex: 150 entries, 0 to 149
## Data columns (total 4 columns):
## sepal length (cm) 150 non-null float64
## sepal width (cm) 150 non-null float64
## petal length (cm) 150 non-null float64
## petal width (cm) 150 non-null float64
## dtypes: float64(4)
## memory usage: 4.8 KB
## Noneg. Subsetting dataframes
# R
#To subset a dataframe 'df' in R we use df[row,column] or df[row vector,column vector]
#df[row,column]
iris[3,4]## [1] 0.2#df[row vector, column vector]
iris[2:5,1:3]## lengthOfSepal widthOfSepal lengthOfPetal
## 2 4.9 3.0 1.4
## 3 4.7 3.2 1.3
## 4 4.6 3.1 1.5
## 5 5.0 3.6 1.4#If we omit the row vector, then it implies all rows or if we omit the column vector
# then implies all columns for that row
iris[2:5,]## lengthOfSepal widthOfSepal lengthOfPetal widthOfPetal Species
## 2 4.9 3.0 1.4 0.2 setosa
## 3 4.7 3.2 1.3 0.2 setosa
## 4 4.6 3.1 1.5 0.2 setosa
## 5 5.0 3.6 1.4 0.2 setosa# In R we can all specific columns by column names
iris$Sepal.Length[2:5]## NULL#Python
# To select an entire row we use .iloc. The index can be used with the ':'. If
# .iloc[start row: end row]. If start row is omitted then it implies the beginning of
# data frame, if end row is omitted then it implies all rows till end
#Python
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
print(iris.iloc[3])
print(iris[:5])
# In python we can select columns by column name as follows
print(iris['sepal length (cm)'][2:6])
#If you want to select more than 2 columns then you must use the double '[[]]' since the
# index is a list itself
print(iris[['sepal length (cm)','sepal width (cm)']][4:7])## sepal length (cm) 4.6
## sepal width (cm) 3.1
## petal length (cm) 1.5
## petal width (cm) 0.2
## Name: 3, dtype: float64
## sepal length (cm) sepal width (cm) petal length (cm) petal width (cm)
## 0 5.1 3.5 1.4 0.2
## 1 4.9 3.0 1.4 0.2
## 2 4.7 3.2 1.3 0.2
## 3 4.6 3.1 1.5 0.2
## 4 5.0 3.6 1.4 0.2
## 2 4.7
## 3 4.6
## 4 5.0
## 5 5.4
## Name: sepal length (cm), dtype: float64
## sepal length (cm) sepal width (cm)
## 4 5.0 3.6
## 5 5.4 3.9
## 6 4.6 3.4h. Computing Mean, Standard deviation
#R
#Mean
mean(iris$lengthOfSepal)## [1] 5.843333#Standard deviation
sd(iris$widthOfSepal)## [1] 0.4358663#Python
#Mean
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
# Convert to Pandas dataframe
print(iris['sepal length (cm)'].mean())
#Standard deviation
print(iris['sepal width (cm)'].std())## 5.843333333333335
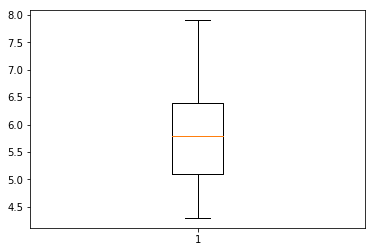
## 0.4335943113621737i. Boxplot
Boxplot can be produced in R using baseplot
#R
boxplot(iris$lengthOfSepal)Matplotlib is a popular package in Python for plots
#Python
import sklearn as sklearn
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
img=plt.boxplot(iris['sepal length (cm)'])
plt.show(img)j.Scatter plot
#R
plot(iris$widthOfSepal,iris$lengthOfSepal)#Python
import matplotlib.pyplot as plt
import sklearn as sklearn
import pandas as pd
from sklearn import datasets
data = datasets.load_iris()
# Convert to Pandas dataframe
iris = pd.DataFrame(data.data, columns=data.feature_names)
img=plt.scatter(iris['sepal width (cm)'],iris['sepal length (cm)'])
#plt.show(img)k. Read from csv file
#R
tendulkar= read.csv("tendulkar.csv",stringsAsFactors = FALSE,na.strings=c(NA,"-"))
#Dimensions of dataframe
dim(tendulkar)## [1] 347 13names(tendulkar)## [1] "X" "Runs" "Mins" "BF" "X4s"
## [6] "X6s" "SR" "Pos" "Dismissal" "Inns"
## [11] "Opposition" "Ground" "Start.Date"Use pandas.read_csv() for Python
#Python
import pandas as pd
#Read csv
tendulkar= pd.read_csv("tendulkar.csv",na_values=["-"])
print(tendulkar.shape)
print(tendulkar.columns)## (347, 13)
## Index(['Unnamed: 0', 'Runs', 'Mins', 'BF', '4s', '6s', 'SR', 'Pos',
## 'Dismissal', 'Inns', 'Opposition', 'Ground', 'Start Date'],
## dtype='object')l. Clean the dataframe in R and Python.
The following steps are done for R and Python
1.Remove rows with ‘DNB’
2.Remove rows with ‘TDNB’
3.Remove rows with absent
4.Remove the “*” indicating not out
5.Remove incomplete rows with NA for R or NaN in Python
6.Do a scatter plot
#R
# Remove rows with 'DNB'
a <- tendulkar$Runs != "DNB"
tendulkar <- tendulkar[a,]
dim(tendulkar)## [1] 330 13# Remove rows with 'TDNB'
b <- tendulkar$Runs != "TDNB"
tendulkar <- tendulkar[b,]
# Remove rows with absent
c <- tendulkar$Runs != "absent"
tendulkar <- tendulkar[c,]
dim(tendulkar)## [1] 329 13# Remove the "* indicating not out
tendulkar$Runs <- as.numeric(gsub("\\*","",tendulkar$Runs))
dim(tendulkar)## [1] 329 13# Select only complete rows - complete.cases()
c <- complete.cases(tendulkar)
#Subset the rows which are complete
tendulkar <- tendulkar[c,]
dim(tendulkar)## [1] 327 13# Do some base plotting - Scatter plot
plot(tendulkar$BF,tendulkar$Runs)#Python
import pandas as pd
import matplotlib.pyplot as plt
#Read csv
tendulkar= pd.read_csv("tendulkar.csv",na_values=["-"])
print(tendulkar.shape)
# Remove rows with 'DNB'
a=tendulkar.Runs !="DNB"
tendulkar=tendulkar[a]
print(tendulkar.shape)
# Remove rows with 'TDNB'
b=tendulkar.Runs !="TDNB"
tendulkar=tendulkar[b]
print(tendulkar.shape)
# Remove rows with absent
c= tendulkar.Runs != "absent"
tendulkar=tendulkar[c]
print(tendulkar.shape)
# Remove the "* indicating not out
tendulkar.Runs= tendulkar.Runs.str.replace(r"[*]","")
#Select only complete rows - dropna()
tendulkar=tendulkar.dropna()
print(tendulkar.shape)
tendulkar.Runs = tendulkar.Runs.astype(int)
tendulkar.BF = tendulkar.BF.astype(int)
#Scatter plot
plt.scatter(tendulkar.BF,tendulkar.Runs)## (347, 13)
## (330, 13)
## (329, 13)
## (329, 13)
## (327, 13)m.Chaining operations on dataframes
To chain a set of operations we need to use an R package like dplyr. Pandas does this The following operations are done on tendulkar data frame by dplyr for R and Pandas for Python below
- Group by ground
- Compute average runs in each ground
- Arrange in descending order
#R
library(dplyr)
tendulkar1 <- tendulkar %>% group_by(Ground) %>% summarise(meanRuns= mean(Runs)) %>%
arrange(desc(meanRuns))
head(tendulkar1,10)## # A tibble: 10 × 2
## Ground meanRuns
##
## 1 Multan 194.00000
## 2 Leeds 193.00000
## 3 Colombo (RPS) 143.00000
## 4 Lucknow 142.00000
## 5 Dhaka 132.75000
## 6 Manchester 93.50000
## 7 Sydney 87.22222
## 8 Bloemfontein 85.00000
## 9 Georgetown 81.00000
## 10 Colombo (SSC) 77.55556#Python
import pandas as pd
#Read csv
tendulkar= pd.read_csv("tendulkar.csv",na_values=["-"])
print(tendulkar.shape)
# Remove rows with 'DNB'
a=tendulkar.Runs !="DNB"
tendulkar=tendulkar[a]
# Remove rows with 'TDNB'
b=tendulkar.Runs !="TDNB"
tendulkar=tendulkar[b]
# Remove rows with absent
c= tendulkar.Runs != "absent"
tendulkar=tendulkar[c]
# Remove the "* indicating not out
tendulkar.Runs= tendulkar.Runs.str.replace(r"[*]","")
#Select only complete rows - dropna()
tendulkar=tendulkar.dropna()
tendulkar.Runs = tendulkar.Runs.astype(int)
tendulkar.BF = tendulkar.BF.astype(int)
tendulkar1= tendulkar.groupby('Ground').mean()['Runs'].sort_values(ascending=False)
print(tendulkar1.head(10))## (347, 13)
## Ground
## Multan 194.000000
## Leeds 193.000000
## Colombo (RPS) 143.000000
## Lucknow 142.000000
## Dhaka 132.750000
## Manchester 93.500000
## Sydney 87.222222
## Bloemfontein 85.000000
## Georgetown 81.000000
## Colombo (SSC) 77.555556
## Name: Runs, dtype: float649. Functions
product <- function(a,b){
c<- a*b
c
}
product(5,7)## [1] 35def product(a,b):
c = a*b
return c
print(product(5,7))
## 35
Conclusion
Personally, I took to R, much like a ‘duck takes to water’. I found the R syntax very simple and mostly intuitive. R packages like dplyr, ggplot2, reshape2, make the language quite irrestible. R is weakly typed and has only numeric and character types as opposed to the full fledged data types in Python.
Python, has too many bells and whistles, which can be a little bewildering to the novice. It is possible that they may be useful as one becomes more experienced with the language. Also I found that installing Python packages sometimes gives errors with Python versions 2.7 or 3.6. This will leave you scrambling to google to find how to fix these problems. These can be quite frustrating. R on the other hand makes installing R packages a breeze.
Anyway, this is my current opinion, and like all opinions, may change in the course of time. Let’s see!
I may write a follow up post with more advanced features of R and Python. So do keep checking! Long live R! Viva la Python!
Note: This post was created using RStudio’s RMarkdown which allows you to embed R and Python code snippets. It works perfectly, except that matplotlib’s pyplot does not display.
Also see
1. My book ‘Deep Learning from first principles:Second Edition’ now on Amazon
2. Dabbling with Wiener filter using OpenCV
3. My book ‘Practical Machine Learning in R and Python: Third edition’ on Amazon
4. Design Principles of Scalable, Distributed Systems
5. Re-introducing cricketr! : An R package to analyze performances of cricketers
6. Natural language processing: What would Shakespeare say?
7. Brewing a potion with Bluemix, PostgreSQL, Node.js in the cloud
8. Simulating an Edge Shape in Android
To see all posts click Index of posts